Meet a scientist: Dr. Linyi Gao breaks down CRISPR genome editing technology
CRISPR technology is one of the major bioengineering breakthroughs of the 21st century. You might have heard all about its potential in editing DNA and curing genetic diseases—or even performed it yourself with our Knockout! or Chopped! Learning Labs. But scientists are continuing to improve this technology, and now, there’s even more to learn about this powerful and fascinating system.
To tell us a bit more about CRISPR, we interviewed Dr. Linyi Gao. Dr. Linyi Gao completed his PhD in biological engineering at MIT in Feng Zhang’s lab, working on microbiology and CRISPR genome editing. He is currently a Junior Fellow with the Society of Fellows of Harvard University.
CRISPR/Cas is able to target and cut specific DNA sequences, which allows for those sequences to be edited or changed. Can you summarize how the targeting and cutting elements of the CRISPR/Cas system work and explain why CRISPR/Cas was so revolutionary?
CRISPR is naturally found in microbes—including the bacteria in yogurt—and exists to help the microbes fight off viruses. CRISPR consists of specialized proteins that team up with guide RNAs to form molecular scissors. These protein-RNA complexes have two jobs to do: find a target site in DNA that matches the guide RNA, and cut the target. The “magic” of CRISPR lies in the fact that the guide RNA can easily be changed to almost anything we want, which allows us to repurpose CRISPR to target specific locations in our own genome.
The human genome is a sequence of about 3 billion letters of DNA, consisting of As, Ts, Cs, and Gs in a defined order. Printed out in small font, it would fill an entire bookcase. Finding a single location within this sequence is a big challenge, analogous to finding a needle in a haystack. The fact that CRISPR can do this with ease underlies the power of the technology.
In addition to editing the human genome, CRISPR can also be used to edit the genomes of animals, plants, and microbes, leading to many other transformative applications throughout basic research, biotechnology, agriculture, and medicine.
How did you become interested in working with this technology?
When I started graduate school, I was excited by all the new possibilities that genome editing enabled, to accelerate basic research and for therapeutics, and I wanted to contribute to the development of the technology. In the following years, I did my PhD mentored by Dr. Feng Zhang at MIT and the Broad Institute, which was a terrific experience not only because of the exciting science but also because of the chance to work with some amazing colleagues.
Can you explain one of the CRISPR projects you’ve worked on before?
CRISPR isn’t perfect. Sometimes it will also edit unintended locations scattered across the genome (off-target editing) just because they happen to have a sequence similar to the guide. For basic research, usually this isn’t a big issue, but when we’re talking about using CRISPR to treat genetic diseases, it’s essential to minimize off-target edits, as they could lead to cancer and other serious consequences if they happen in the wrong genes.
In 2015, a colleague and I explored whether the inherent precision of the Cas9 protein could be improved by tweaking its amino acid sequence. Surprisingly, we found that changing even a single amino acid (out of 1,368 total), if done in the right place in the protein, could dramatically reduce the amount of off-target editing while keeping about the same level of on-target activity.
When we look at a CRISPR/Cas system, the two main components are the Cas nuclease and the guide RNA. Cas9 is likely the one that people have heard of, but there are many different types of Cas nucleases out there. Can you explain the differences between the commonly used Cas nucleases?
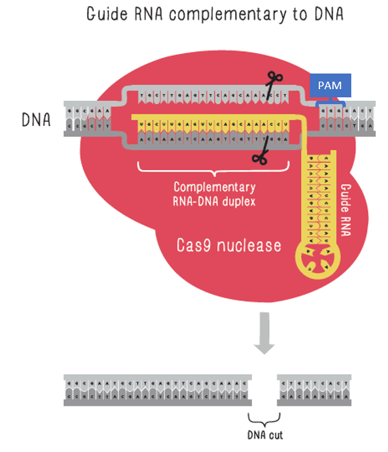
Cas9 is the nuclease most commonly used for genome editing. Besides the gRNA matching to the target DNA, the Cas9 also needs to match its PAM, a three-base sequence of NGG, where N can be any base, near the target DNA. If both these conditions are met, the Cas9 will cut through both strands of the DNA.
Besides Cas9, two other types of commonly used CRISPR nucleases exist: Cas12 and Cas13. Cas12a has a shorter guide RNA and has a different PAM that contains mostly T’s instead of G’s. On the other hand, Cas13 proteins don’t cleave DNA at all, but rather RNA. These different properties allow them to be used in applications like CRISPR-based diagnostics, instead of genome editing.
It’s worth noting that CRISPR systems in nature are remarkably diverse, and there are many more flavors of CRISPR that exist in microbes than the versions we like to use for genome editing. Some of these are useful in their own right—for example, by using smaller proteins or different PAMs.
One of the powerful aspects of CRISPR/Cas is how guide RNAs can be designed by just using base-pairing rules, but sometimes it takes a few tries to get the best guide RNA for your experiment. What are scientists currently doing to improve guide RNA designs?
Picking a good guide RNA is key for any genome engineering experiment. Different target sequences can differ by an order of magnitude or more in terms of how effectively Cas enzymes can act on them, as well as in the number of unintended off-target sites.
Since there are hundreds of millions of possible CRISPR target sites in the human genome, it isn’t feasible to just experimentally test them all beforehand and be done with it. Instead, the most effective approach for guide RNA prediction has been to develop computational models that can predict good guide RNAs based on experimental data. These data have typically been collected from pooled screens that simultaneously assess the performance of thousands of guides or more. The resulting models have become state-of-the-art tools that are now widely used to assist in improved guide RNA design.
CRISPR biology remains a rich and active area of research, with hidden gems still being uncovered in many different kinds of microbes. Nature is an amazing artist!
Are there any other developments in the CRISPR/Cas system that excite you?
Recent engineering innovations have expanded the capabilities of the CRISPR toolbox beyond DNA cleavage. For example, by combining a partially-inactivated Cas9 with other DNA-modifying enzymes unrelated to CRISPR, researchers have been able to more precisely change the target DNA sequence without actually cleaving it. Known as base editors and prime editors, these engineered fusion proteins represent the next generation of CRISPR technology.
In addition, CRISPR biology remains a rich and active area of research, with hidden gems still being uncovered in many different kinds of microbes. Nature is an amazing artist!
Which potential application of CRISPR is the most exciting to you?
I’m excited about the potential of CRISPR to treat or cure a subset of genetic diseases. A number of clinical trials have already seen promising results, such as for sickle cell disease, with surely more on the way in the coming years. It’s inspiring to see CRISPR-based technologies move from bacteria all the way to the clinic and have a profound positive impact on people’s lives.
Do you have any advice for students who are interested in a career in science?
I think it’s valuable to get exposure to diverse scientific fields in college and before. As an undergrad, I took classes in biology, chemistry, physics, and applied math, which I’ve found helpful in my career as a scientist. Grad school is a good time to dive deep into a research topic, but because the day-to-day of bench science can be very time-consuming, it becomes harder (though not impossible) to find the bandwidth to systematically learn things outside your research field.
This interview by Dr. Ally Huang has been edited for length and clarity.
Related resources:
- Student questions for Dr. Gao interview
- CRISPR Teaching Resources
- Video: Genome editing with CRISPR/Cas9
- Article: First patients to get CRISPR gene-editing treatment continue to thrive
- Research article: Rationally engineered Cas9 nucleases with improved specificity (by Dr. Linyi Gao and colleagues)