Meet a Scientist: Jon Hale studies genetic diversity in daffodils
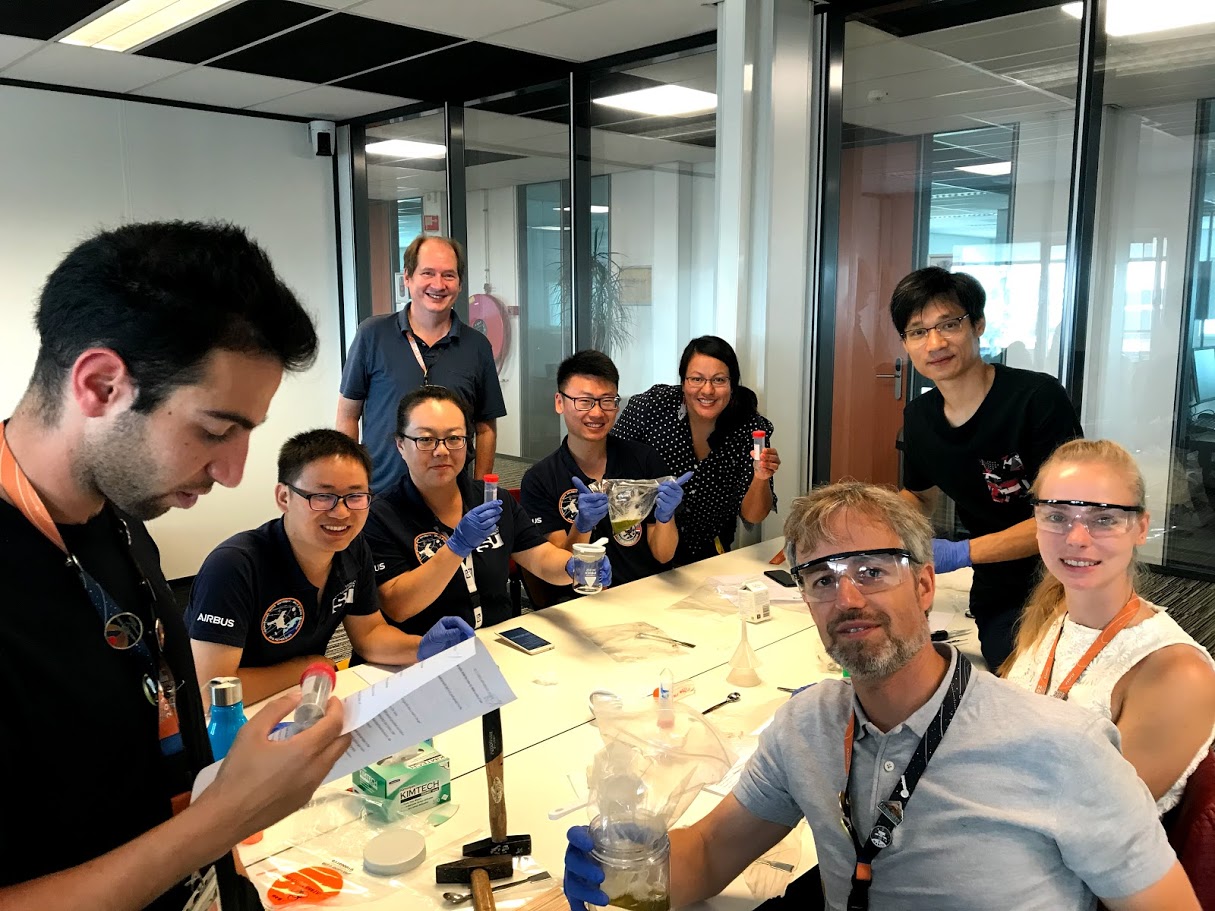
Led by teacher Jon Hale on the Island of Jersey (UK), the Daffodil DNA Project brings together scientists from universities and research institutes with schools to sequence the chloroplast genomes of different varieties (cultivars) of daffodils. The scientific goal of this project is to obtain genetic data on this very understudied, yet valuable genus. To learn more about this exciting work, we interviewed Mr. Jon Hale, Head of Biology at Beaulieu Convent School in Jersey, UK.
What is the Daffodil DNA Project?
When you think of genetics you might remember Gregor Mendel and his investigations hybridizing pea plants which he shared in 1865. Perhaps you might link the name William Bateson who shared Mendel’s findings with researchers in England in 1902, but what you might not know is that these people were not geneticists, but instead interested in flowers, and how to breed the best ones.
We have learned so much about genetics from plants, like the “jumping genes” discovered in maize by Barbara McClintock, however we have shifted our focus towards human genetics, or genetics of human pathogens like the coronavirus in recent years. There are still scientists working on some crop species, such as wheat and barley, and the model organism Arabidopsis thaliana, but this is a drop in the ocean in terms of what knowledge is still possible to discover. The Daffodil DNA Project aims to do just that, add to scientific knowledge of an under-researched, yet still important, plant.
The Daffodil DNA project brings together scientists from universities and research institutes with schools to sequence the chloroplast genomes of different varieties (cultivars) of daffodils.
Some of the daffodils we have sequenced are historically important, having been bred before the 1900s and are grown in very low numbers, usually within the national collections.
How did you become interested in daffodils and how did the project get started?
The project was inspired by a springtime stroll in the countryside, taking the time to notice nature and observe that not all daffodils are yellow. There were loads of different colors, some had ruffled cups, some had big trumpets, some were huge, some were tiny. It turns out that selective breeding has led to over 30,000 named cultivars.
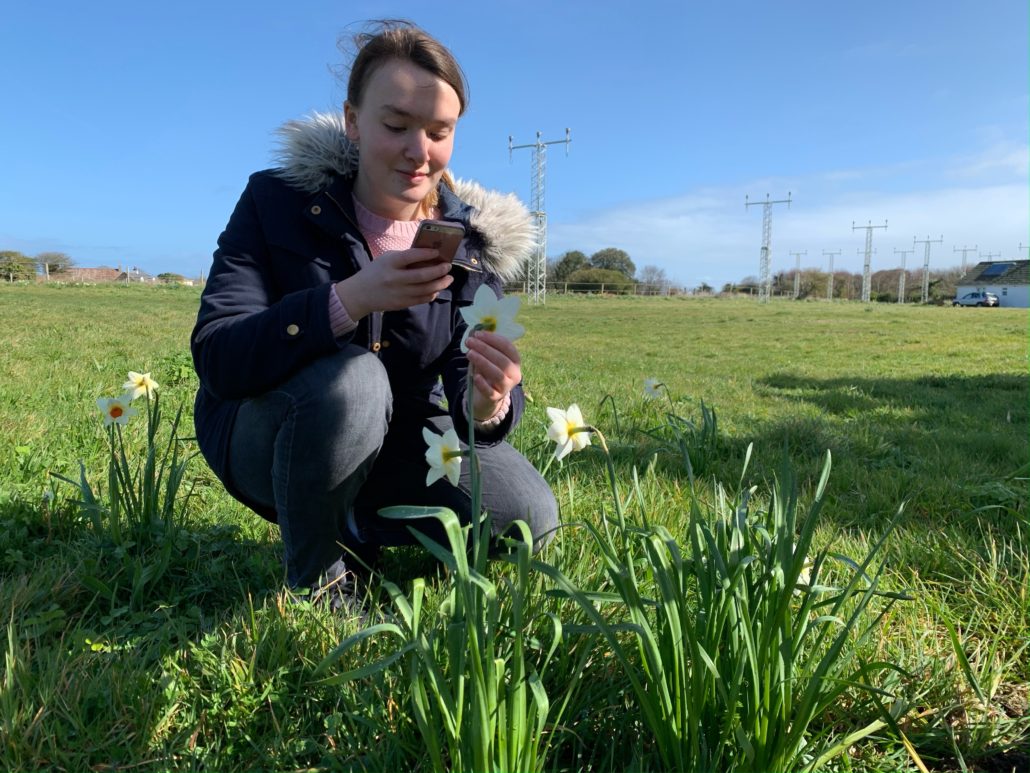 The biology of daffodils is very cool. They produce toxic molecules called alkaloids. Some alkaloids, like narciclasine, can kill other flowers. Others, like lycorine, affect animals, causing severe vomiting and diarrhea if eaten. There’s a reason why cows and sheep don’t eat daffodils in their fields. There is also an alkaloid produced by daffodils called galantamine which has been used to treat the symptoms of Alzheimer’s.
The biology of daffodils is very cool. They produce toxic molecules called alkaloids. Some alkaloids, like narciclasine, can kill other flowers. Others, like lycorine, affect animals, causing severe vomiting and diarrhea if eaten. There’s a reason why cows and sheep don’t eat daffodils in their fields. There is also an alkaloid produced by daffodils called galantamine which has been used to treat the symptoms of Alzheimer’s.
Despite the daffodil’s interesting metabolism, the biochemical research into daffodil DNA is not vast. We are far away from a complete genome, but there have been three complete chloroplast genomes for different daffodils published. The ability to stand on the shoulders of giants has unlocked this project.
Schools use cutting edge technology to generate sequencing data by preparing the DNA library using a miniPCR thermocycler before loading nanopore flow cells.
These flow cells detect the tiniest changes in the DNA molecules’ characteristics as they pass through the pores before computers interpret the signals as base sequences, those As, Ts, Cs and Gs. We then align each sequenced fragment against the published chloroplast genome sequences to build our own sequences. When we’ve got these sequences, we can then compare the similarities to try and work out if they are closely related, or not.
What is special about chloroplast genomes?
The common view is that chloroplasts are descended from a bacterium that smuggled its way into eukaryotic cells. Their ability to photosynthesize lead to a mutually beneficial relationship with the host cell, allowing it to thrive and reproduce. Over time, lots of the genes within the chloroplast genome moved into the nuclear genome, but lots still remain. Structurally, most of it seems to be in a loop, like a bacterial genome, but some people have seen branching and linear forms which might explain some of our data anomalies too.
How are students involved in the project?
Through the collection and analysis of DNA samples, students aim to identify and map the genetic traits that give rise to different daffodil flower colors, shapes, sizes, and other features. This information can then be used to better understand daffodil genetics, as well as to develop new cultivars with desirable traits.
Students learn to extract DNA from daffodil leaves and use high throughput DNA sequencing in the classroom before assembling the chloroplast’s genome. In this process they use cutting edge genetic technologies: preparing DNA libraries using a miniPCR thermocycler, validation using blueGel electrophoresis, and generating nanopore sequencing data. But the goals of this project extend beyond the purely scientific, as students work with STEM professionals and academic researchers to forge ongoing relationships.
Can you briefly describe the workflow and the tools needed to carry it out?
The first step in the procedure is to smash up the leaves as much as possible using a chilled pestle and mortar. From this green mush, DNA can be extracted using an “off-the-shelf” kit from Qiagen. This washes the DNA and digests the RNA that would be present. Then the DNA is prepared for sequencing by fragmenting the molecules and sticking them to adapter sequences using a thermocycler. The DNA library is then loaded into an Oxford Nanopore minION sequencer.
Once the data has been collected, we carry out a parallel approach to the dry-lab. In the classroom, students use a piece of software called Guppy which introduces them to command line computing before then using a drag-and-drop program called Geneious Prime which they build their assemblies and phylogenetic trees. At the University of Dundee, the computational biologists take the raw files that the students generate and do the same steps using open source software, trying to get the data as good as it can be. The results are simply amazing.
How many schools are now involved? How do they share their data?
There have now been 10 schools involved in the project. Schools are able to upload their data to a shared folder, allowing higher quality basecalling and chloroplast genome assembly using cloud computational power in contrast to the PCs in the classroom.
 What were the biggest challenges setting up the project, and how did you overcome them?
What were the biggest challenges setting up the project, and how did you overcome them?
In the UK biology classroom, we all share two challenges: time and money. Getting micropipettes, microcentrifuges, thermocyclers and DNA sequencers is a considerable financial burden for most schools. However there are some very generous and supportive grant schemes. We applied for a Royal Society Partnership Grant, which provides up to £3,000 for resources. This was more than enough to make the project come to fruition and be financially sustainable.
Time, however, is much harder to come by. It is a finite commodity, but it can be maximized through creativity. The daffodil is not a model organism for our teaching of much of our post-16 curriculum. This means that our sequencing is embedded, rather than a bolt-on. This has allowed our project to evolve from an extracurricular opportunity to a key part of every biology student’s learning experience.
In other schools that have adopted the project, they complete all of the wet-lab activities in one day, taking students off-timetable to allow them to discuss with scientists their career aspirations, run gels and quantify their DNA extractions using blueGel electrophoresis as a bonus.
 Can you tell us more about the people working on the project?
Can you tell us more about the people working on the project?
There are so many heroes involved in the project. There are amazing high school students beginning their journey in a scientific career, being supported by fantastic biology teachers from across the UK. Each school is partnered with an actively researching scientist from the James Hutton Institute, the University of Dundee and now Harper Adams University and the University of West England. These professional scientists go into the school as collaborators, sharing their experience in this field outside of their own research goals.
How can I get involved?
It would be amazing to develop this project far and wide and give more schools and colleges the opportunity to highlight science as a viable career for their students. If you are interested in taking part, please do get in touch and we’ll see how we can start the project in your school or college.
Related resources:
- University of Dundee’s Daffodil DNA Project page.
- Daffodil DNA Project on Twitter
- Daffodil DNA Project on YouTube
- DNAdots: Nanopore sequencing