Taking Mendel Molecular with PCR, Gel Electrophoresis and Wisconsin Fast Plants®
Wisconsin Fast Plants® have been a staple of Biology classrooms for decades. With short generation times, high productivity, and many distinct visual phenotypes, generations of students have used Fast Plants® to study a variety of topics ranging from Mendelian genetics, to artificial selection, to energy flow through ecosystems. But as molecular techniques such as PCR and gel electrophoresis have entered high school classrooms, a limitation of Fast Plants® has been the limited availability of genetic sequence information. This has left a gap for teachers who would like to link classical genetics breeding with molecular genetics approaches.
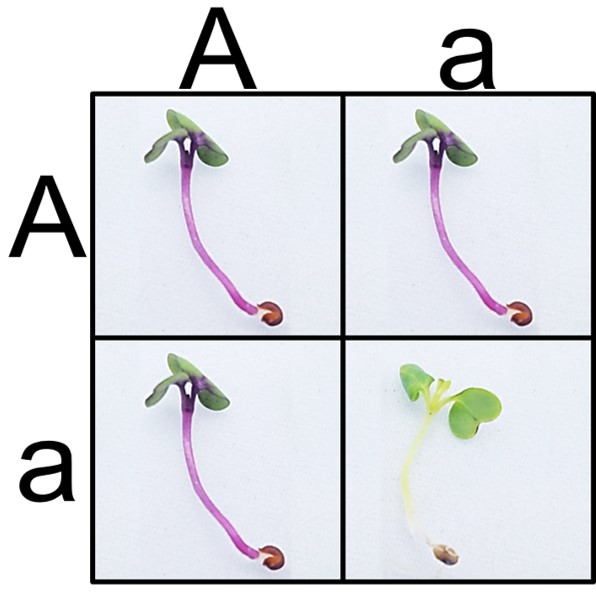
The miniPCR™ Plant Genetics Lab looks to bridge that gap. Students and teachers can now link a clearly observable phenotype to its underlying genetic cause. Purple versus non-purple stem color is a commonly used trait in classrooms, especially when investigating Mendelian genetics. Based on recently published research by Wendell et al. [1], we now know that Anthocyaninless, the gene responsible for the two phenotypes, is a gene that codes for the production of the enzyme Dihydroflavonol 4-reductase (DFR), an integral enzyme in the production of the purple pigment, anthocyanin.  The difference in the two alleles is a 354 base pair insertion in the recessive green allele (anl, or a) that knocks out the function of the resulting protein. The dominant purple allele (ANL, or A) produces a functional DFR protein. Using PCR and gel electrophoresis, students can now amplify the Anthocyaninless gene to determine the genotype of their plants based on this insertion.
The difference in the two alleles is a 354 base pair insertion in the recessive green allele (anl, or a) that knocks out the function of the resulting protein. The dominant purple allele (ANL, or A) produces a functional DFR protein. Using PCR and gel electrophoresis, students can now amplify the Anthocyaninless gene to determine the genotype of their plants based on this insertion.
A classic and quick approach using Fast Plants® in the classroom is to purchase F2 seeds derived from an original AA x aa parental cross and to germinate them. Students can then see if seeds match the expected 3:1 phenotypic ratio according to Mendelian predictions and test their data using the Chi-squared statistic. With the miniPCR Plant Genetics Lab, students can take the approach one step further. Students can now test whether plants also adhere to the 1:2:1 genotypic ratio as predicted by Mendel, all with a simple DNA extraction, PCR, and gel electrophoresis protocol that can be easily completed in two class periods.
 An exciting aspect of this lab is that it can be employed using several different approaches. Wisconsin Fast Plants® are designed to have short generation times with prolific seed production, so several generations can be bred by students in a class over a few weeks. This allows teachers to take a more inquiry, student-centered approach. Students can grow the parental strains and create their own F1 and F2 plants. Students can then predict the genotypic and phenotypic ratios of each generation. By testing the genotypes of parental, F1, and F2 plants, students can actually visualize the inheritance of alleles over time.
An exciting aspect of this lab is that it can be employed using several different approaches. Wisconsin Fast Plants® are designed to have short generation times with prolific seed production, so several generations can be bred by students in a class over a few weeks. This allows teachers to take a more inquiry, student-centered approach. Students can grow the parental strains and create their own F1 and F2 plants. Students can then predict the genotypic and phenotypic ratios of each generation. By testing the genotypes of parental, F1, and F2 plants, students can actually visualize the inheritance of alleles over time.
Some teachers take the student-centered inquiry approach even further. Students are given seeds of unknown genotype and asked to cross them and germinate the offspring. Then by observing the phenotypes of the offspring they must predict the genotypes of the parents. With the miniPCR Plant Genetics Lab, students can save samples from the parental plants and confirm if their analysis of the offspring was correct – effectively testing their predictions and seeing real and authentic applications of biotechnology in the classroom.
The miniPCR machine was invented as an affordable biotechnology tool that could put real science in the hands of students, increasing both access and engagement. We are excited to introduce this new lab, extending already engaging classroom activities in ways that were not previously possible. Being able to show that molecular techniques are not a separate endeavor, but an extension to the science that students are already doing is an invaluable way of connecting advanced techniques to basic biology.
Join the DNA revolution!
Fast Plants® is a registered trademark of the Wisconsin Alumni Research Foundation.
[1] Wendell, D.L. Vaziri, A. Shergill, G. (2016) The Gene Encoding Dihydroflavonol 4-Reductase Is a Candidate for the anthocyaninless Locus of Rapid Cycling Brassica rapa (Fast Plants Type) PLoS One, 11(8)