From Snakes to Sequences in 24 Hours: Success!
— This case study contributed by Aaron Pomerantz and Stefan Prost from the Ecuadorian rain forest appeared originally on 7/19/2017 in thenextgenscientist.com. It illustrates that in times when we are losing the natural world faster than ever, using new tools can make biology and conservation more efficient. Rapid DNA sequencing in the field can be a useful tool to better understand the diversity of life on our planet and use that information for conservation.
— 11/17/2017 update: Watch this great video summarizing the rainforest expedition by Aaron Pomerantz and Stefan Prost
As I type up my notes from the field, I can only imagine what Darwin would think about the advancements in technology used to answer questions about evolutionary biology today…
7/11/17 6:00 AM
We hit the ground running after sharing the last post at the airport. Landing at around 6:00 AM in Quito, Ecuador, we were greeted by our collaborators Lucas Bustamante (Tropical Herping) and Dr. David Salazar (Universidad Indoamerica), threw our gear in the truck, and swung by the University lab to pick up our final crew member, Nicolás Peñafiel. All the equipment and reagents for sequencing appeared to have traveled well on ice packs (at least we hoped so!), so we transferred our frozen material into a cooler and were off to the Chocó rainforest of Ecuador.

7/11/17 3:00 PM
After several hours of driving through bumpy jungle roads and crossing the Canandé river on a questionable boat, we arrived at the small lodge nestled in the Chocó rainforest and rendezvoused with the rest of the team, who arrived a couple days ahead of us, including Alejandro Arteaga, Frank Pichardo, and Cesar Barrio Amorós.
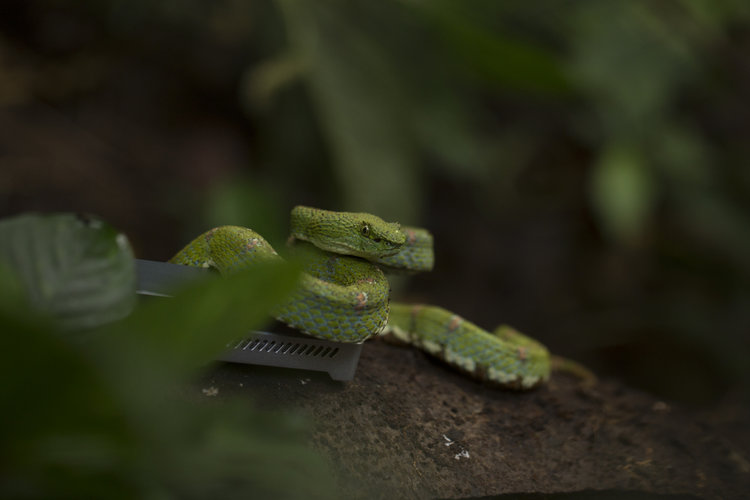
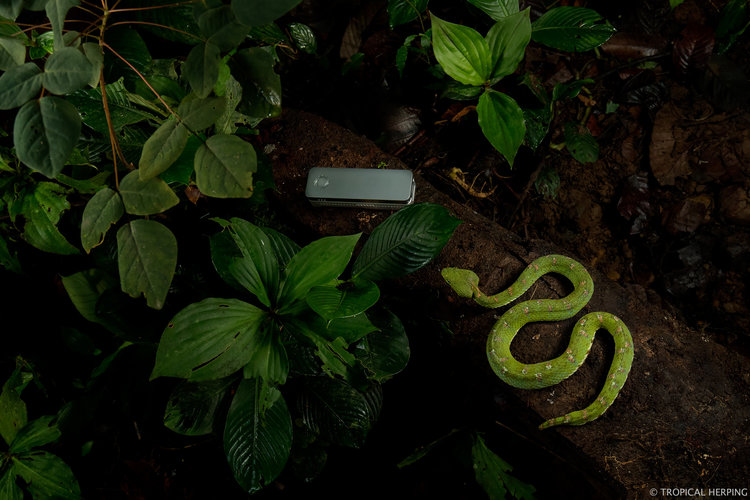
Our goal was to survey the region for unique reptiles and amphibians and use the portable laboratory to sequence DNA right there in the field, so no time to waste! Nico and I unpacked the lab equipment and at around 8:24 PM began a few DNA extractions on a whip snake sample using a salt extraction protocol and the DNeasy kit protocol. Then as night fell, we grabbed our headlamps and were out in search of new targets. After several hours spotting numerous frogs and geckos, Frank stumbled upon a gorgeous eyelash viper just off trail.


7/11/17 11:34 PM
Back at the lodge that evening, we prepped a PCR run using the miniPCR and let it go overnight. Time for some rest.
7/12/17 10:15 AM
While Alejandro took digital white background images of the animals from last night, Nico and I made a gel to visualize the PCR product. The small electrophoresis chamber Nico brought from the lab worked well, but unfortunately my small UV flashlight was unable to really pick up fluorescence in the gel to visualize amplicons. I had tested this out briefly in the US before the trip and was able to pick up bands in a pitch black room, but I think the UV light wasn’t quite strong enough under field conditions. Some of the bands appeared to be there which was encouraging, but next time I think it would be beneficial to tinker with a small transilluminator like the one also made by miniPCR.
7/12/17 11:30 AM
Alejandro wrapped up processing his specimens and extracting a small amount of blood or tail tissue from each, so I got to work extracting DNA for the eyelash viper and dwarf geckos. The extraction process with the DNeasy kit is easy (as the name implies!); as for equipment just requires a small centrifuge and the reagents can be stored at room temperature. After about an hour we had our fresh DNA samples.
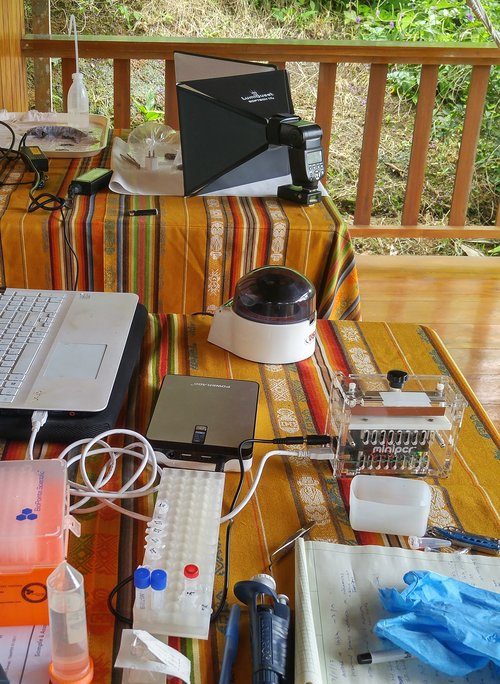

Next I got started with a new round of PCR using primers for genes for 16S, cytb and ND4. These primers don’t all necessarily have the same PCR conditions (such as annealing temperature) but for the sake of time and limitations with one miniPCR, I ran them together under the same settings.
3:25 PM
After a couple hours of PCR cycles, it was time for the second PCR barcding step. I used barcodes 1 through 8 for the samples and ran the new PCR protocol.
4:30 PM
Finally, it was time to start the library preparation for the nanopore sequencer using the SQK-LSK 1D kit. This involves the end-prep, adapter ligation and bead cleanup. Next it was time to prime the flow cell and load the sample.
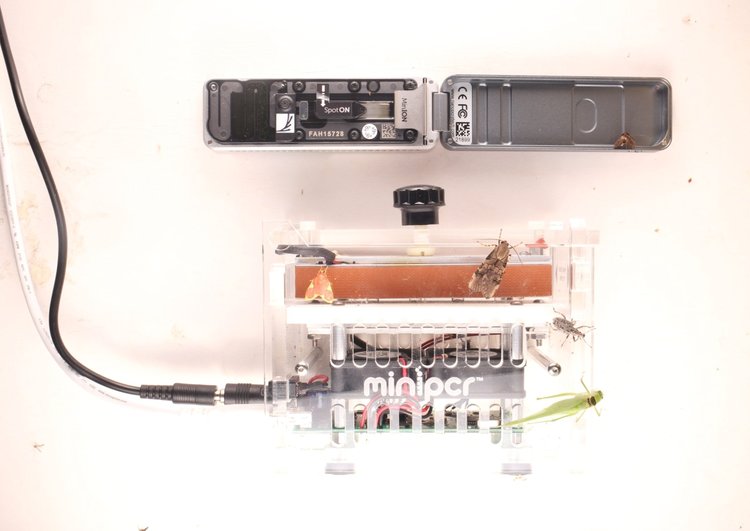

6:24 PM
I clicked “execute” for the MinKNOW software, began the sequencing run and said ‘hold on to your butts’ (one of my favorite quotes from Jurassic Park).


7:20 PM
After about an hour, I stopped the run after 16,484 reads had been generated, and ran the data through the Albacore program to demultiplex the reads into their individual barcode folders. I then took a peak in the barcodes and was excited to see the read lengths looked correct, so I downloaded barcode 1 and passed it along to Alejandro’s laptop. Barcode 1 was the 16S sequence for the eyelash viper and Alejandro had a nice reference database on his laptop to compare the nanopore sequence to. After a few minutes of tinkering, Ale said “yes, it falls out with Bothriechis schlegelii!“. This was it! The nanopore barcode was a complete match to the correct species!


In less than 24 hours from arriving at the field site and sampling snakes and geckos during our first night in the rainforest, we could verify species identification by creating a nanopore consensus and mapping to a pre-downloaded reference database. Now we are also verifying if some species collected are undescribed back at a lab in Quito, and will have everything Sanger sequenced to verify nanopore quality.
9:30 PM
After getting back from the field, Stefan set out to process the reads. While Geneious is good enough for a quick peek, it cannot deal with Nanopore-specific errors. After some quick tests using reference-based mapping (using bwa mem, samtools, angsd and nanopolish) it was clear we got good consensus sequences for all the 16S and the ND4 genes! However, CytB and COI did not amplify, which we verified on a gel back at Quito university a few days later, likely because we didn’t have time to run multiple PCRs under optimal contitions for all the genes. Stefan then worked on tweaking de-novo assembly tools such as Canu and Allele Wrangler, to create consensus sequences without reference bias. All sequences from the field were later processed with Canu, which turned out to work pretty well for amplicon assembly.
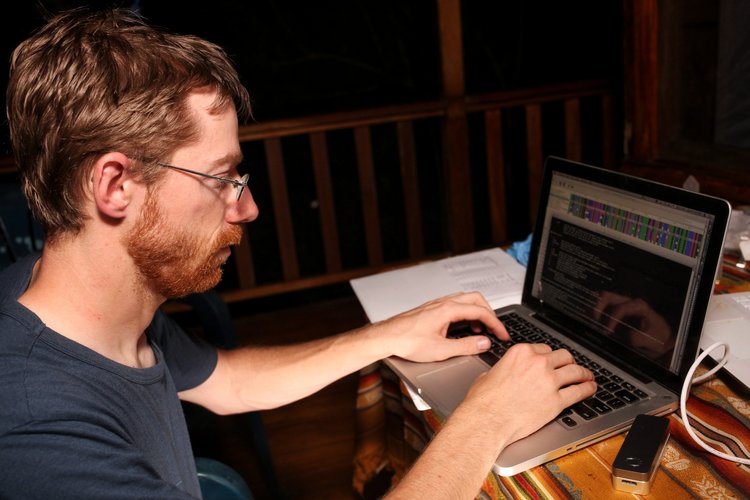

Overall, we believe this is important becuase the Ecuadorian Chocó is a biodiversity hotspot which has lost more than 98% of habitat due in large part to logging and palm oil agriculture. Rapid sequencing can be a useful tool to better understand the diversity of life on our planet and use that information for conservation. Furthermore, researchers in Quito do not have access to a Sanger sequencer let alone Next Generation Sequencing platforms within the country. Thus, Oxford Nanopore MinION sequencing enables them to rapidly process samples without the need to send them off internationally.

Right now I’m still feeling a bit of relief and excitement that everything worked in one go in the field. This was an idea that I’ve been hoping to execute for a while now, and this seemed like the opportune time with a grant funding the project from National Geographic. The work isn’t quite done yet, because we are verifying the quality of sequences, performing a few more experiments in lab, and writing all the methods and bioinformatics pipelines up for a publication with our Ecuadorian collaborators. While we quickly and successfully sequenced DNA in the field for correct species identification, this feels like just the beginning!