Studying Genomes in the Madagascar Forest
Whether due to bureaucratic red tape or the remoteness of the site, field researchers have long been plagued by the problem of how to bring the samples they collect back to a place where they can study them. Elaine Guevara was no exception to this rule. As a PhD candidate in biological anthropology at Yale, she made regular trips to the dry, spiny forests of Madagascar to visit a population of wild lemurs that has been the subject of long-term research by multiple groups, including her own. In particular, Guevara focused on the population’s genetic history and diversity, which she studied by capturing the animals and taking cheek swabs once in their lifetimes —a frequency (or, rather, infrequency) that made each sample precious.

But the colony of sifaka lemurs Guevara studied at Beza Mahafaly Reserve in Southwest Madagascar is endangered, and the logistical gymnastics required to transport biological matter belonging to such a species were complex. By the time researchers were able to remove the samples from the distant island nation and send them to a lab where Guevara would be able to extract and amplify the DNA, the swabs had already spent long periods stored under less-than-optimal conditions, jeopardizing both their own quality and the analysis that could be performed on them.
Faced with such an obstacle, Guevara considered an unusual alternative. If the difficulty lay in bringing the samples to the lab, she asked, why not bring the lab to the samples? She had heard from a colleague about a portable thermal cycler produced by the biotech startup Amplyus, and it seemed promising as a solution to her problem—a machine that she could carry into the field to amplify DNA on the spot. Powering the miniPCR through a lithium ion battery that she charged from a small solar panel, Guevara was able to begin performing whole genome amplification (WGA) assays straight from the remote Madagascar field site.

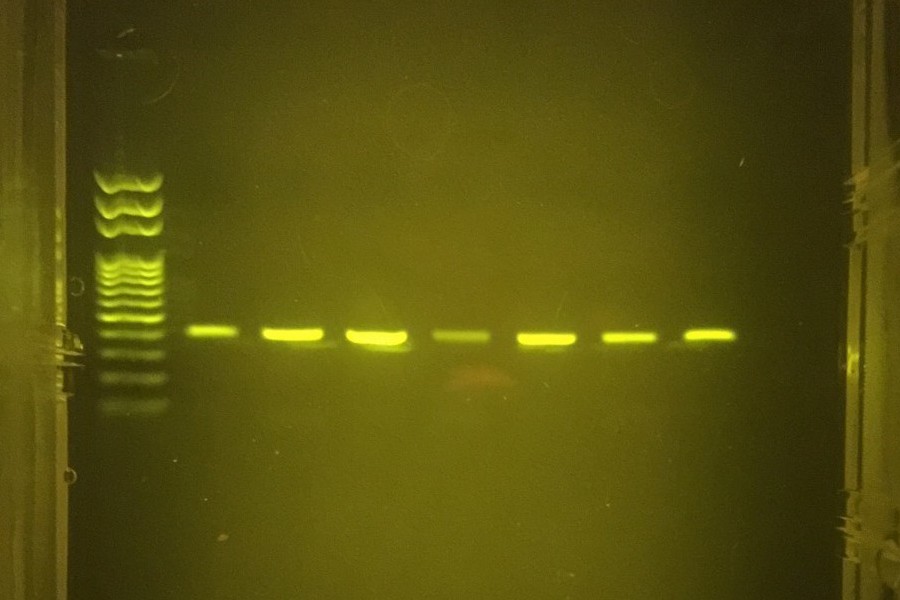
Only one question now remained: would the WGAs work? That is, would the miniPCR be able to stand up to the task at hand as well as the bench-top machines upon which the team had previously relied? The answer, Guevara asserts, is yes. “The miniPCR seems just as effective as a traditional thermocycler, but of course far more transportable, durable, and energy-efficient,” she says. It would seem that with the miniPCR, Guevara had found a way to overcome the challenge that had for so long stymied her peers—and the resulting enhancements in the condition of the DNA and the extent of analysis that could be conducted on it, of course, speak for themselves.
Today, having returned from Madagascar after a first round of gene analysis, Guevara is excited for the horizons that have opened to her research, and she hopes to continue to use the miniPCR to facilitate her findings. “The use of the miniPCR represented a major improvement for us because we were able to obtain and amplify DNA right there in the field,” she says. “Going forward, I am hoping the miniPCR will also allow us to do even more of the genetic analyses in Madagascar…. I think miniPCR is ideal for the field.”
Elaine’s microsatellite genotyping of these lemurs will help determine relatedness for the population, and can answer questions about mating strategies, fitness, and dispersal, as well as help reconstruct the demographic history of the population and assess their genetic diversity.
Download this case study by clicking on this link.
Read the peer reviewed publication this study led to here.
– Victoria Lin
Related resources:
Elaine Guevara is a Ph.D. candidate in Biological Anthropology, interested in molecular evolution and the genetic basis of primate traits. Visit her webpage at Yale University Department of Anthropology, and read more about her research here.