miniPCR™ Training for Space on the Ocean Floor
This guest blog post was contributed by Sarah Stahl, M.S., and Sarah Wallace, Ph.D., spaceflight microbiologists at NASA.
Microbiological monitoring is an essential activity on the International Space Station (ISS). Microbiologists at the Johnson Space Center in Houston, TX, monitor the air, water, and surfaces of the ISS for microbial contamination to ensure a safe habitat for the crew. Currently on the ISS, technologies capable of microbial identification and infectious disease diagnosis are not available. Meeting these needs relies on a time- and resource-intensive process that involves collection and culture of microbial samples on orbit by the astronauts and sample return for ground-based identification by DNA sequencing. With the technological development of portable, miniature, molecular biology tools, the potential to implement DNA sequencing in space has become a reality.
The Biomolecule Sequencer payload, launching to the ISS on SpaceX-9, will assess the functionality of the commercially-available MinION™ (Oxford Nanopore Technologies) in the microgravity environment of space. For this experiment, ground-prepared samples containing bacterial, viral, and murine genomic DNA will be launched and stored frozen on the ISS until sequencing initiation. Successful completion of the spaceflight experiment will demonstrate, for the first time, that DNA sequencing in space is possible. As DNA sequencing is a tool that can be used beyond microbial identification, this validation will open the doors to far wider potential, such as providing near real-time information on scientific investigations, monitoring cellular changes in response to spaceflight, informing medical interventions, defining the effectiveness of countermeasures, and aiding in the detection of nucleic acid-based life elsewhere in the universe. However, this next level of application potential will not be feasible until an end-to-end, sample collection-to-data analysis process is developed.
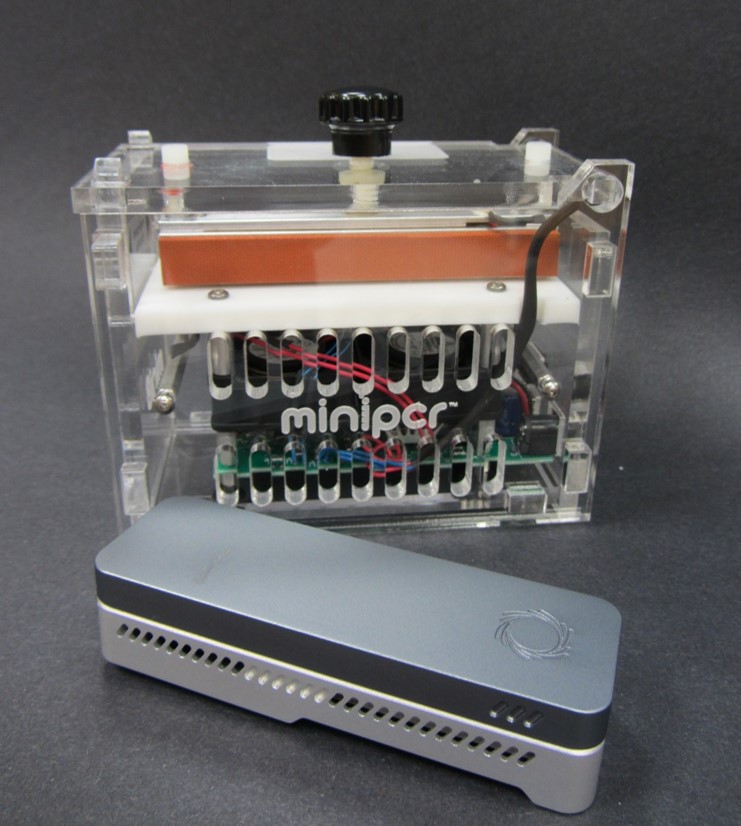

The Biomolecule Sequencer payload team is working toward the development of a spaceflight-compatible swab-to-sequencer sample preparation process. Historical data has demonstrated that microbes are in low abundance in the air, water, and on surfaces of the ISS. As such, it is anticipated that low levels of DNA will be generated from the extraction of these low biomass samples. To overcome this hurdle, an amplification step will be inserted between the DNA extraction and library preparation components of the protocol. The amplification of DNA is simplistic in theory and practice, yet traditional thermalcyclers used to perform this reaction are large and possess less than ideal power requirements for spaceflight. The miniPCR™ system is an exception, as it is small, simplistic in design, has a high ease-of-use, can be powered by a laptop or tablet, and, most importantly, is spaceflight approved and already aboard the ISS after its successful launch on SpaceX-8 in April 2016. As part of its first mission, miniPCR is being used to test whether the polymerase chain reaction (PCR) can be used to study DNA alterations aboard the ISS. These reasons make miniPCR™ a logical choice for inclusion in a sample preparation process for DNA sequencing in space. The entire swab-to-sequencer process will be spaceflight analog tested during the NASA Extreme Environment Mission Operations (NEEMO) 21 mission in July, 2016.
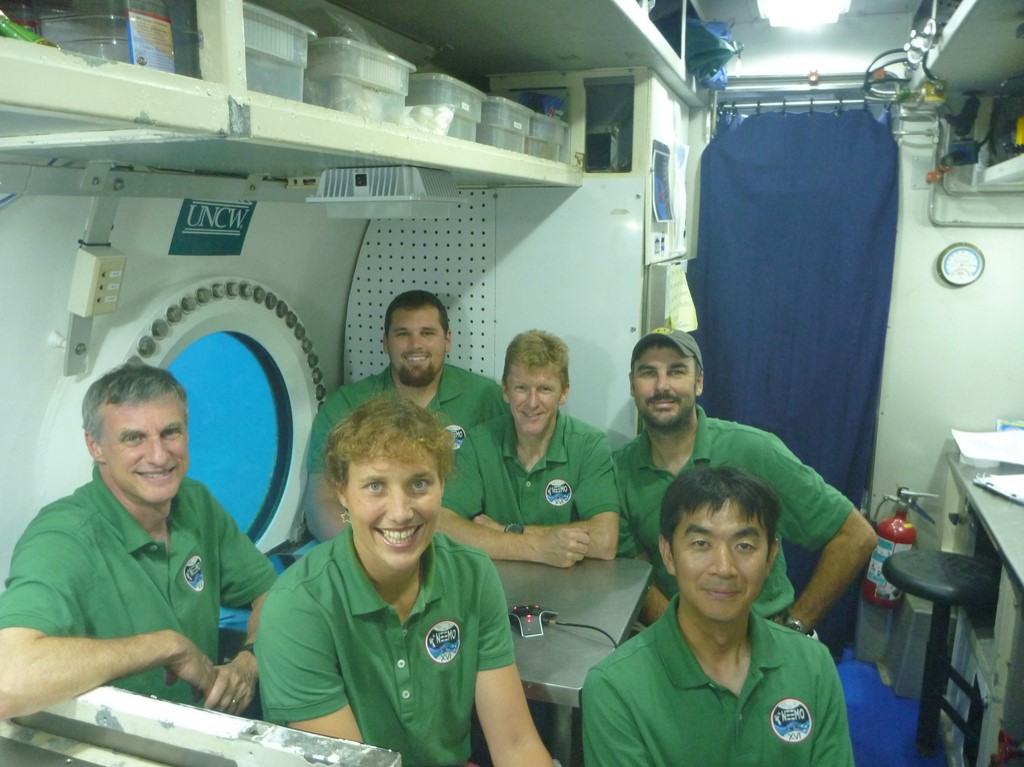
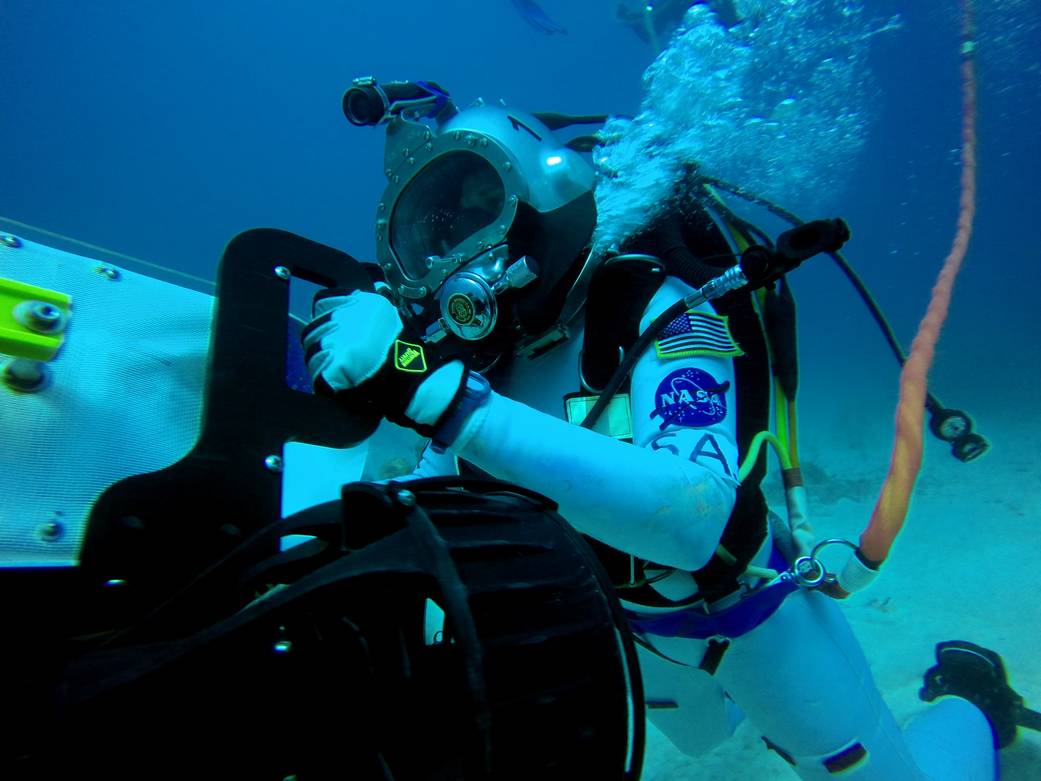
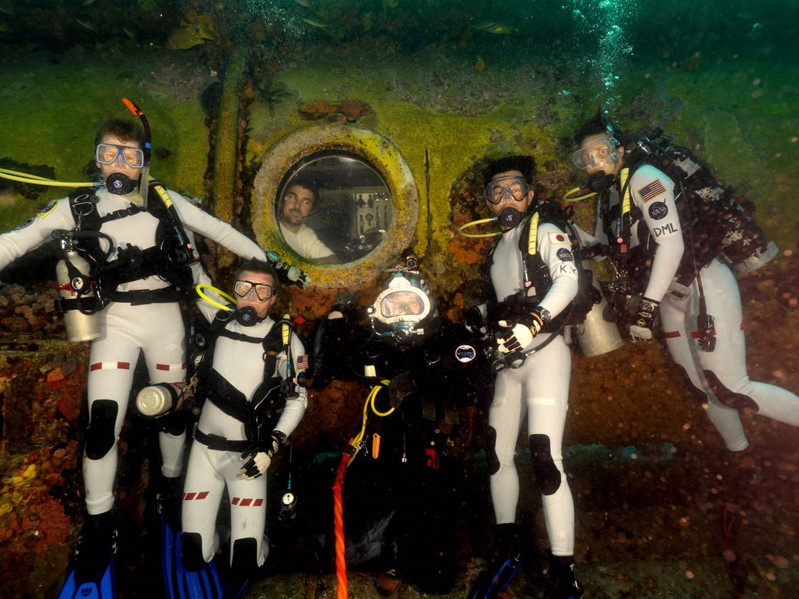
The NEEMO habitat, Aquarius, is the world’s only underwater research station and is located on the ocean floor approximately five miles off the coast of Key Largo, FL, at a depth of 62 feet. Due to its extreme location and the challenges presented, NASA’s NEEMO missions serve as an analog to spaceflight missions where astronauts, scientists, and engineers live and work to test new hardware and solve spaceflight-related problems.
Images of NEEMO and NEEMO crews (Credit: NASA)