Fighting Superbugs with AI: A New Hope in Antibiotic Discovery
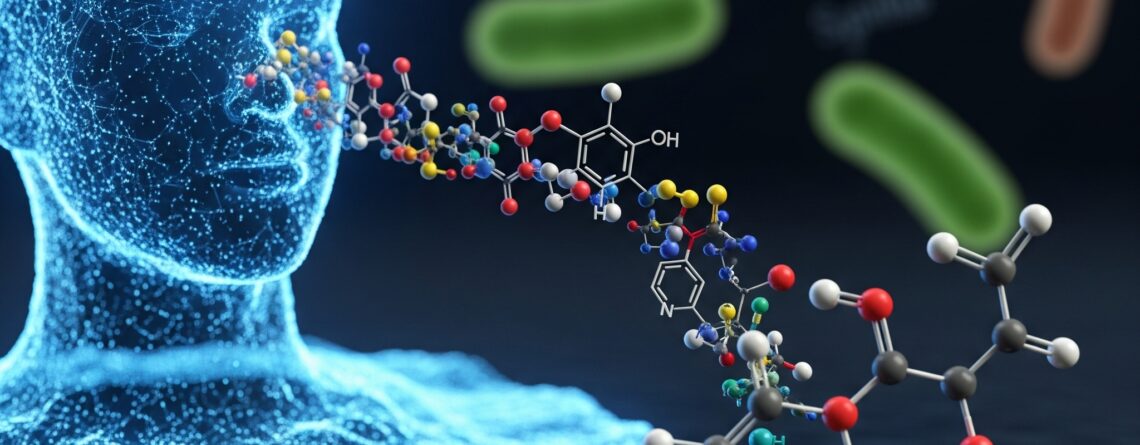
Antibiotic resistance is one of the most pressing public health crises of our time, responsible for millions of deaths worldwide. For decades, the pace of new antibiotic development has slowed dramatically, leaving us with a dwindling arsenal against a growing threat. Researchers at MIT and Harvard, in a study published in Cell, are tackling this challenge head-on by using a powerful new tool: generative artificial intelligence (AI). This groundbreaking approach doesn’t just screen existing compounds; it designs entirely new ones from scratch, opening up a vast, uncharted chemical landscape in the search for effective drugs.
Think of it like this: traditional drug discovery is a bit like searching for a needle in a haystack—a very, very big haystack of chemical compounds. AI-powered virtual screening can make this search faster and more efficient, but it’s still limited by what’s already in the haystack. The new approach is a game-changer. It’s like teaching a machine how to grow new hay to make new, better needles.
 The research team developed a powerful AI platform that uses two main methods to generate new molecules. The first, a fragment-based approach, starts with small chemical building blocks and expands on them. The team screened over 45 million chemical fragments to find those predicted to have antibacterial activity against two dangerous bacteria: Neisseria gonorrhoeae and Staphylococcus aureus. Once promising fragments were identified, two generative AI models—a genetic algorithm called CReM and a variational autoencoder (VAE)—were used to design complete, drug-like molecules that incorporated these fragments.
The research team developed a powerful AI platform that uses two main methods to generate new molecules. The first, a fragment-based approach, starts with small chemical building blocks and expands on them. The team screened over 45 million chemical fragments to find those predicted to have antibacterial activity against two dangerous bacteria: Neisseria gonorrhoeae and Staphylococcus aureus. Once promising fragments were identified, two generative AI models—a genetic algorithm called CReM and a variational autoencoder (VAE)—were used to design complete, drug-like molecules that incorporated these fragments.
The second method is even more ambitious: de novo design. This approach generates entirely new molecules without any starting point, like a blank canvas. The AI models use their learned knowledge of chemistry to create millions of new, unique compounds from basic inputs like water or methane.
The results were astonishing. Out of the thousands of molecules the AI designed, the researchers synthesized and tested a select few. Of the 24 compounds synthesized, seven showed selective antibacterial activity. Two of these, named NG1 and DN1, were particularly impressive.
NG1 was designed using the fragment-based approach and was found to be highly effective against multidrug-resistant strains of N. gonorrhoeae. Its unique mechanism of action involves targeting a protein called LptA, which is essential for building the bacterial outer membrane but is absent in human cells. This unique target helps NG1 bypass existing resistance mechanisms, offering a new way to fight a rapidly evolving threat. NG1 was also effective in a mouse model of infection.
DN1 was a product of the unconstrained de novo design and proved potent against multidrug-resistant S. aureus. Like NG1, its mechanism is also membrane-related, causing morphological changes and disruption of the bacterial membrane. DN1 also showed strong efficacy in a mouse model of skin infection.
This research highlights the incredible potential of generative AI to push the boundaries of drug discovery and address the urgent need for new antibiotics. By designing molecules that explore previously untouched chemical spaces, this platform offers a powerful new strategy in our fight against superbugs.
Classroom activities to understand and address antibiotic resistance
Dive into the mechanisms of antibiotic resistance and raise awareness about this critical public health issue with these hands-on activities, with innovative kits that provide a hands-on approach for students to explore the science behind antibiotic resistance and discover cutting-edge solutions in the fight against superbugs.
- BioBits®: Antibiotic Resistance: This activity allows students to visualize the action of antibiotics on gene expression in a cell-free system. They can directly observe how a genetic sequence can produce a protein that renders an antibiotic ineffective, demonstrating mechanism of antibiotic action and resistance.
- True Blue™ Transformation Lab: Genetically Engineering Bacteria: Students will genetically engineer bacteria to express a specific gene, providing a tangible model for how bacteria can acquire new traits, including antibiotic resistance, through gene transfer.
- Knockout! A CRISPR/Cas Gene Targeting Lab: This lab introduces students to the revolutionary CRISPR/Cas gene-editing technology. They can target and “knock out” a specific gene, offering a powerful tool to understand how scientists can manipulate bacterial genomes and potentially develop new strategies to overcome resistance.
- eDNA Project: Sampling Soil for Antibiotic Resistance: Students use molecular techniques to help monitor the spread of antibiotic resistance in the environment.
These labs don’t just teach the science; they provide a platform to discuss the real-world implications of antibiotic resistance and inspire the next generation of scientists and public health leaders to tackle this global challenge.